Attribute-based interpretable representations of medical images with variational autoencoders
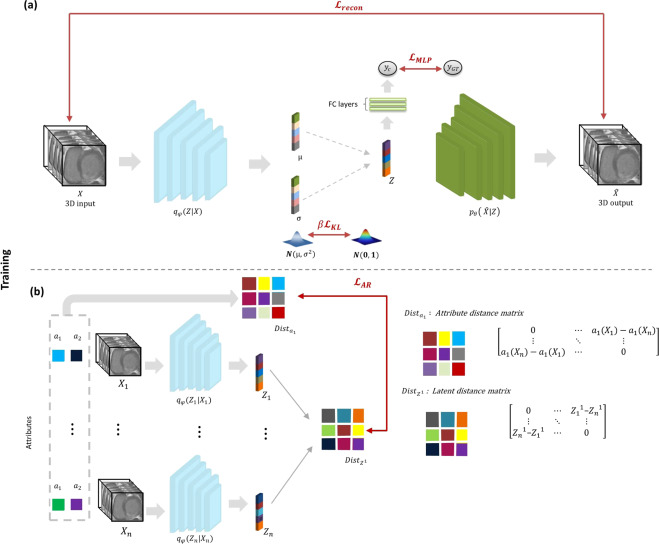
Deep learning (DL) methods are required to better understand the relationship of clinical and imaging-based attributes with DL outcomes. Thus, facilitating their use in the reasoning behind the medical decisions. In this paper, a variational autoencoders (VAE) approach is proposed, the Attri-VAE, that includes an attribute regularization term to associate clinical and medical imaging attributes with different regularized dimensions in the generated latent space, enabling a better interpretation of the attributes. Using the Attri-VAE approach healthy and myocardial infarction patients with clinical, cardiac morphology, and radiomics attributes have been analyzed. The proposed model provided an excellent trade-off between reconstruction fidelity, disentanglement, and interpretability, outperforming state-of-the-art VAE approaches according to several quantitative metrics. The resulting latent space allowed the generation of realistic synthetic data in the trajectory between two distinct input samples or along a specific attribute dimension to better interpret changes between different cardiac conditions.
To read more: https://www.sciencedirect.com/science/article/pii/S0895611122001288

